Our community
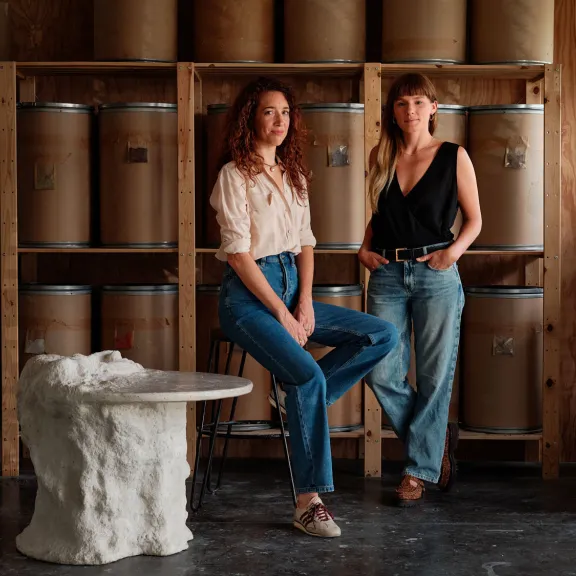
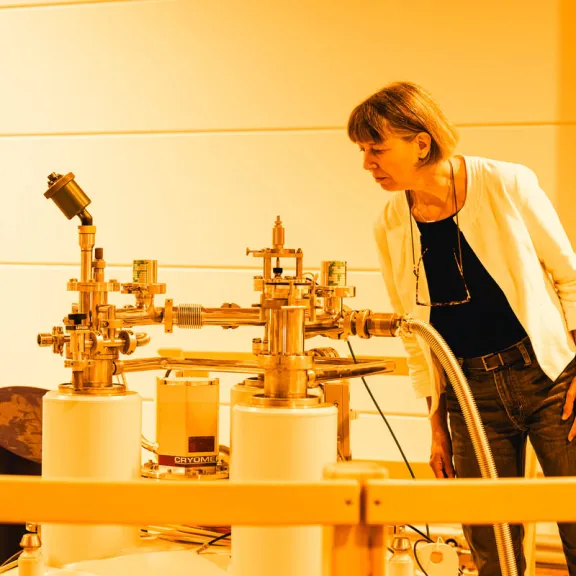
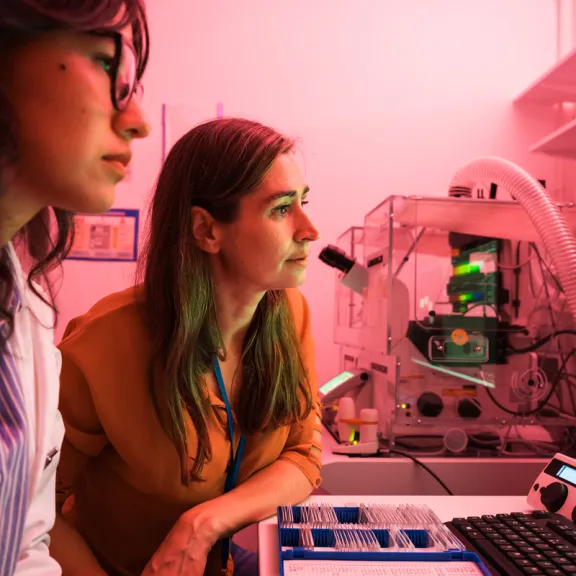
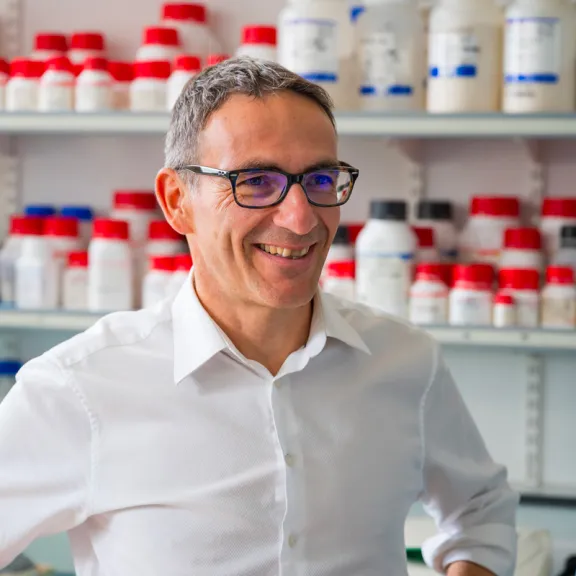
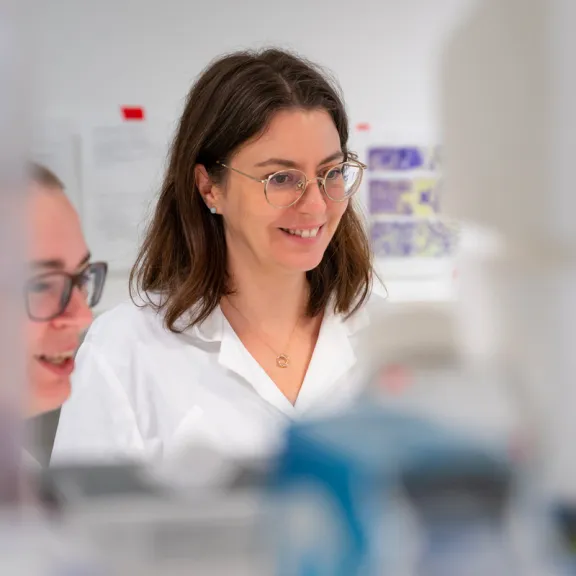
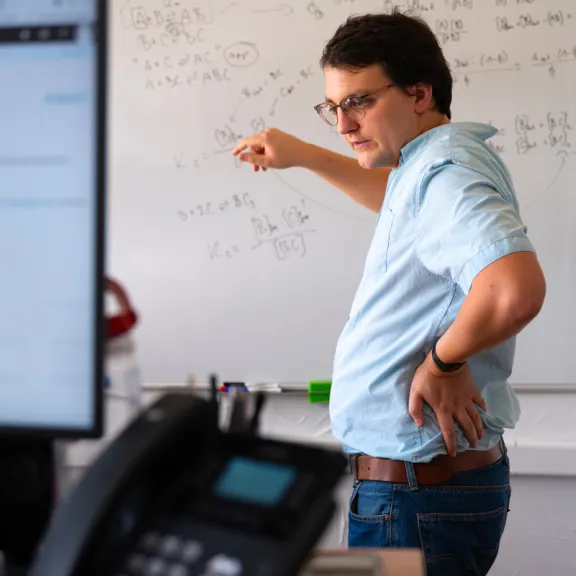
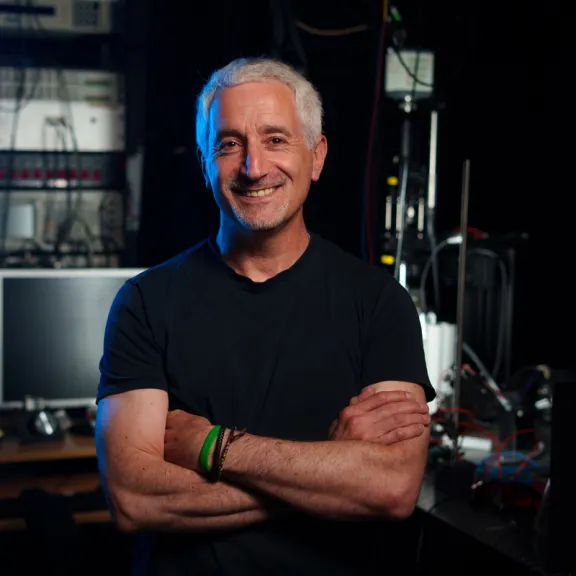
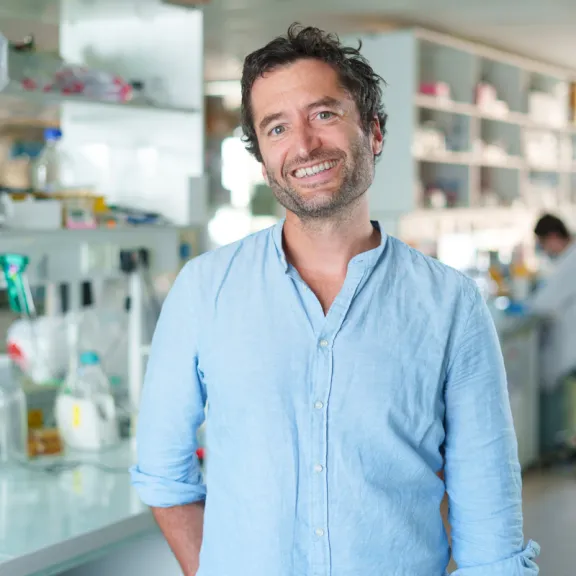
Meet the award-winners and projects leaders supported by the Foundation
Search in our community












Arts - Craftsmanship
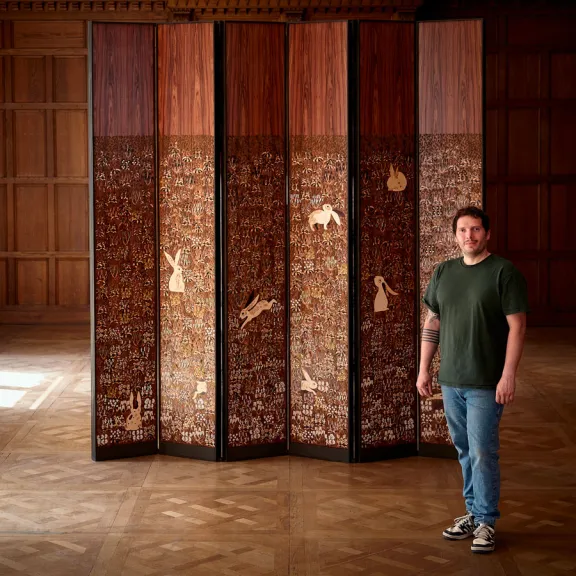
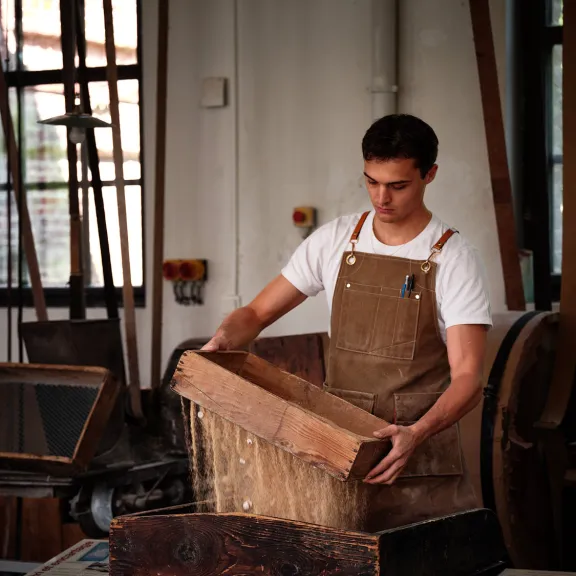
The Musée de la Nacre et de la Tabletterie
Bringing age-old craftsmanship back to life